TL;DR: A 2026 Nature Medicine study found that GBA1 Parkinson’s risk carriers already had gut microbiome shifts before motor symptoms, with about 25% of their signature sitting between healthy controls and diagnosed Parkinson’s patients.
Key Findings
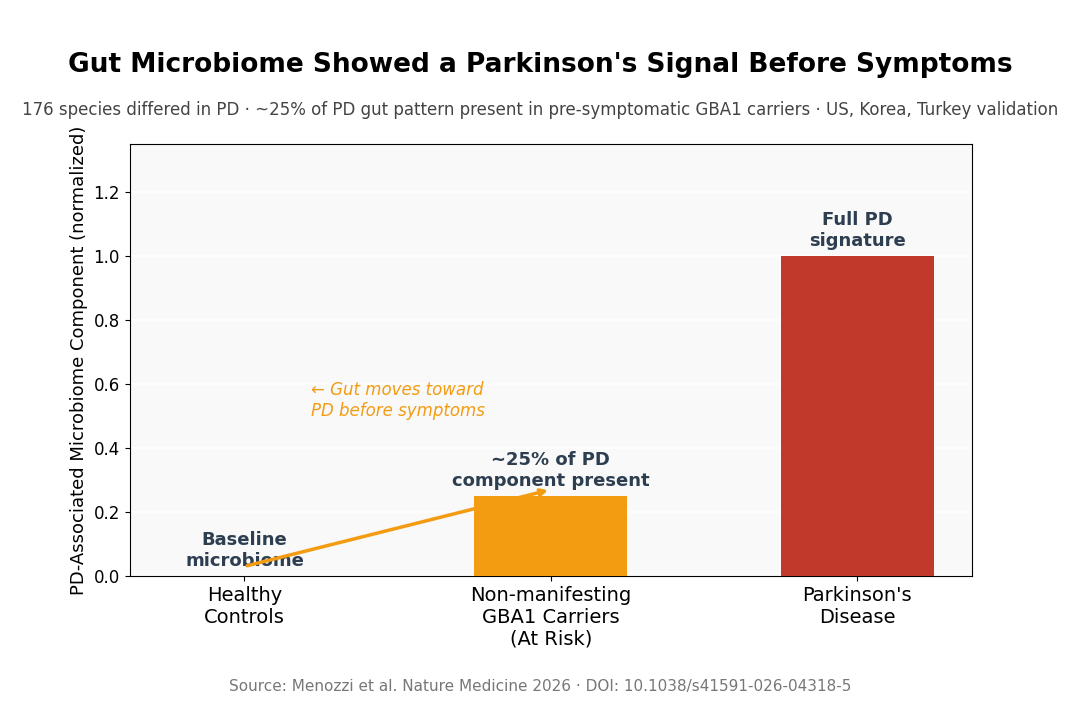
- Pre-symptomatic gut looked intermediate: About 25% of the Parkinson’s-linked microbiome component in non-manifesting GBA1 carriers sat between healthy controls and diagnosed patients — not fully normal, not fully Parkinsonian.
- 176 species differed in Parkinson’s disease: Discovery cohort of 271 PD patients, 43 at-risk GBA1 carriers, and 150 controls; species analysis identified enrichment of Bifidobacterium and depletion of butyrate-producing taxa.
- Prodromal symptoms tracked the gut microbiome pattern: The intermediate microbiome pattern correlated with prodromal features — REM sleep behavior disorder, constipation, hyposmia — in at-risk participants.
- Three-country validation held up: The Parkinson’s-linked microbiome pattern replicated in independent cohorts from the US, Korea, and Turkey (638 additional PD patients, 319 controls).
- The microbiome pattern was not limited to genetic cases: The microbiome signature appeared in non-GBA1-variant Parkinson’s patients too, suggesting it may reflect a broader disease process rather than one genetic subtype.
- Cross-sectional design — conversion not yet proven: The study identifies a pre-symptomatic pattern; it doesn’t yet show which GBA1 carriers will actually develop Parkinson’s or over what timeframe.
Source: Nature Medicine (2026) | Menozzi et al.
Parkinson’s disease has a brutal diagnostic delay built in.
Motor symptoms — the tremor, stiffness, characteristic slowness — typically appear only after 60–80% of dopaminergic neurons in the substantia nigra are already gone. By the time a neurologist confirms the diagnosis, the key biological window for neuroprotection may have long since closed.
This is why the field has been hunting for early biomarkers for decades. And increasingly, those hunts are going somewhere unexpected: the gut.
A Nature Medicine study found that the microbiome of people genetically at risk for Parkinson’s but not yet symptomatic already looked partly Parkinson-like — and the degree of that similarity tracked with how many prodromal warning signs they were carrying.
GBA1 Carriers Are the Right Stress Test for an Early Biomarker
GBA1 is the most common strong genetic risk factor for Parkinson’s disease — mutations in this gene, which encodes a lysosomal enzyme involved in cellular waste clearance, raise lifetime risk by several-fold. But most carriers never develop the disease. That makes non-manifesting GBA1 carriers an unusually useful population: elevated risk, uncertain outcome, no current symptoms.
If a biomarker can distinguish between GBA1 carriers who are already progressing toward Parkinson’s and those who aren’t, it’s offering something much more valuable than a genotype test. The microbiome’s intermediate signature in this paper does exactly that — it reflects where a carrier sits on a disease trajectory, not just whether they carry a risky allele.
The core finding: roughly 25% of the Parkinson’s-linked microbiome component in non-manifesting carriers was intermediate between healthy controls and diagnosed PD patients. The gut had already started moving in the Parkinson’s direction while the clinic still read “well.”
176 Species Built a Pattern, Not a Single Biomarker Bug
The species-level analysis gave the study its biological texture. In the 271 PD patients versus 150 healthy controls, 176 differentially abundant species marked the distinction. The pattern included increased Bifidobacterium and depletion of butyrate-associated organisms in families like Lachnospiraceae and Ruminococcaceae.
Butyrate producers aren’t passive members of the microbiome. They maintain gut barrier integrity, have documented anti-inflammatory properties, and produce a short-chain fatty acid that crosses to the brain via multiple routes. Losing them while gaining taxa that thrive in an inflamed, permeability-disrupted intestinal environment fits the hypothesis that Parkinson’s risk is entangled with gut immune tone long before neurons start dying.
But the paper’s most important framing is that the finding is a pattern, not a single organism. Complex neurodegenerative diseases don’t announce themselves through one heroic bacterium. The diagnostic information lives in a coordinated shift across many species — which is also why the finding needs multi-site validation to survive, not just a replication cohort next door.
The Intermediate Microbiome Pattern Tracked Prodromal Symptoms
What makes this more than a compositional curiosity is that the intermediate microbiome profile in at-risk carriers wasn’t random. It correlated with the prodromal symptoms that clinicians already recognize as early Parkinson’s warning signs: REM sleep behavior disorder, constipation, and hyposmia (reduced sense of smell).
REM sleep behavior disorder, where people physically act out their dreams due to loss of normal muscle atonia, is one of the strongest prodromal predictors of Parkinson’s and related synucleinopathies. A gut signature that tracks with it in pre-symptomatic carriers means the microbiome pattern isn’t drifting randomly — it’s correlating with a biological state that neurologists already take seriously as an early marker.
This alignment between gut and prodromal symptoms is the study’s conceptual core. It moves the gut from “descriptive association” to “candidate early-warning system that could complement existing prodromal markers.”
International Validation Separated the Microbiome Pattern From Lab Artifact
Microbiome studies have a replication crisis problem. Findings routinely fail to survive geography changes, dietary differences, or different sequencing protocols. This paper took that challenge seriously by validating the Parkinson’s-associated microbiome pattern in independent cohorts from the United States, Korea, and Turkey — 638 PD patients and 319 healthy controls across three very different microbiome environments.
Cross-national persistence doesn’t mean every species replicated identically. That would be unrealistic given how strongly geography, diet, and ethnicity shape the microbiome.
What replicated was the broader compositional pattern. That’s the level of replication that matters for a biomarker that needs to work in diverse clinical populations.
The finding that the signature appeared in non-GBA1-variant Parkinson’s patients adds another layer of generalizability. This isn’t a boutique marker for a rare genetic subtype — it may reflect a more universal component of the disease program.
The Practical Payoff Is Trial Enrichment
The field needs to be clear-eyed about what this means and doesn’t mean yet. This is a cross-sectional study.
It captures a snapshot; it doesn’t prove that GBA1 carriers with an intermediate microbiome signature will convert to Parkinson’s, or on what timeline. A prospective conversion study — following carriers forward and asking which ones develop the disease — is the critical next step.
In the meantime, the most practical near-term application is trial enrichment. Disease-modification studies for Parkinson’s have long struggled because patients enter trials after substantial neurodegeneration. A gut-microbiome signature that helps identify people already progressing toward the disease, but not yet diagnosable, could allow trials to intervene much earlier — while the biological window for neuroprotection is still open.
That would be a major clinical shift. The challenge now is longitudinal confirmation and figuring out how the microbiome pattern should be combined with other prodromal markers — seed amplification assays for alpha-synuclein, smell tests, sleep monitoring, blood biomarkers — rather than used in isolation. A combination biomarker approach will almost certainly outperform any single readout for predicting conversion.
What this paper establishes is the gut’s claim to a seat at that table.
Citation: DOI: 10.1038/s41591-026-04318-5. Menozzi et al. Microbiome signature of Parkinson’s disease in healthy and genetically at-risk individuals. Nature Medicine. 2026.
Study Design: Case-control discovery cohort with prospective international validation; fecal metagenomics and clinical data.
Sample Size: Discovery: 464 participants (271 PD, 43 GBA1 carriers, 150 controls). Validation: 957 participants across US, Korea, Turkey.
Key Statistic: ~25% of the PD-linked microbiome component in non-manifesting GBA1 carriers was intermediate between healthy controls and diagnosed PD patients; pattern correlated with prodromal symptoms and replicated across three countries.